Sonication-Assisted Protein Extraction from Tumor Tissue – Protocol
This protocol describes a sonication-assisted protein extraction method for tumor tissue that advances the standard CPTAC urea lysis workflow by adding a controlled probe-sonication step during tissue disruption. Building on the established deep-scale CPTAC proteome and phosphoproteome preparation strategy, this modification improves cell and subcellular structure disruption, reduces sample viscosity, and enhances release of proteins that are typically harder to recover with urea lysis alone, especially membrane-bound and DNA-binding or nucleus-associated proteins. In the underlying study, the sonication-assisted workflow increased detection of both proteins and phosphopeptides while preserving compatibility with the downstream CPTAC-style digestion, TMT labeling, fractionation, phosphopeptide enrichment, and LC-MS/MS analysis pipeline.
Sonication-Assisted Protein Extraction from Tumor Tissue for Deep Proteomic and Phosphoproteomic Analysis
 The following protocol is optimized for extraction of proteins from cryopulverized tumor tissue in 8 M urea lysis buffer: An added probe-type sonication step improves the recovery of difficult protein classes, especially membrane-associated and DNA-/nucleus-associated proteins, before digestion and downstream LC-MS/MS analysis. In the underlying study by Li et al. (2025), adding sonication increased detection of membrane and nucleus-associated proteins and supported deep-scale coverage of >12,000 proteins and >25,000 phosphopeptides under their workflow.
The following protocol is optimized for extraction of proteins from cryopulverized tumor tissue in 8 M urea lysis buffer: An added probe-type sonication step improves the recovery of difficult protein classes, especially membrane-associated and DNA-/nucleus-associated proteins, before digestion and downstream LC-MS/MS analysis. In the underlying study by Li et al. (2025), adding sonication increased detection of membrane and nucleus-associated proteins and supported deep-scale coverage of >12,000 proteins and >25,000 phosphopeptides under their workflow.
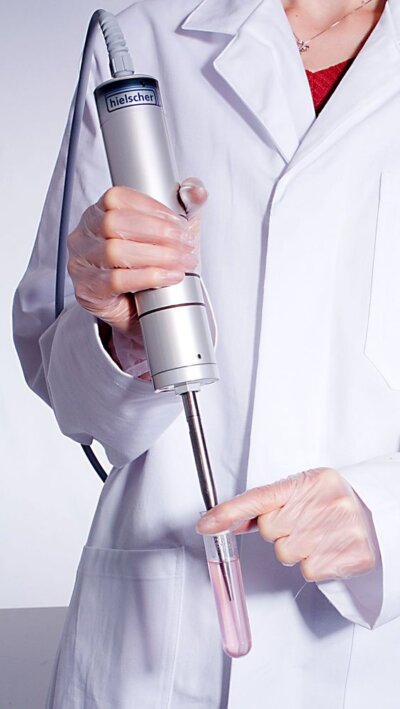

Ultrasonic probe UP200St for protein extraction in proteomics
Application Area for the Protocol
Use this procedure for:
- fresh frozen, cryopulverized tumor tissue
- cell pellets or other biologic specimens where urea-based extraction is already established
- global proteomics and phosphoproteomics workflows using tryptic digestion and optional TMT labeling
The study by Li et al. (2025) states the workflow is also applicable to other specimen types such as cell lines, blood, and urine, however optimization may be required by sample type.
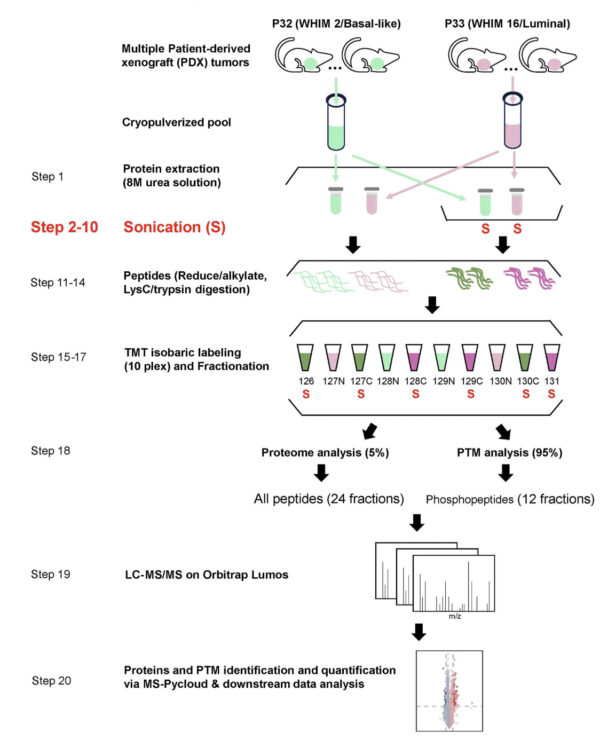
Optimized sample preparation workflow with implemented sonication step. The optimized workflow and experimental design are based on the CPTAC sample preparation protocol for global proteomic and phosphoproteomic analysis.
Study and scheme: ©Li et al., 2025
Working Principle
The original study added sonication to the standard CPTAC urea lysis workflow and found improved detection of proteins associated with membranes and nuclei. The authors state that sonication parameters must be fine-tuned regarding sample size/concentration – by choosing sonotrode size, energy input, and pulse timing.
Exemplary, for the Hielscher UP200Ht and UP200St, this means:
- use amplitude and pulse mode as the main control variables
- keep samples cold throughout (i.e. on ice)
- begin with conservative settings
- optimize against sample clarity, temperature, protein yield, and downstream peptide quality
Both Hielscher 200 watts sonicator models UP200Ht and UP200St are designed for small and medium sample volumes, with adjustable amplitude and pulse settings; both ultrasonic homogenizers provide digital touchscreen control for precise parameter adjustment, automated data recording, pluggable temperature sensor, remote control, sample illumination.
Reagents for Sonication-Assisted Protein Extraction
Urea lysis buffer
Prepare fresh immediately before use:
- 8 M urea
- 75 mM NaCl
- 50 mM Tris, pH 8.0
- 1 mM EDTA
- 2 µg/mL aprotinin
- 10 µg/mL leupeptin
- 1 mM PMSF
- 10 mM NaF
- 20 µM PUGNAc
- Phosphatase Inhibitor Cocktail 2, 1:100 (v/v)
- Phosphatase Inhibitor Cocktail 3, 1:100 (v/v)
Urea must be completely dissolved before adding additives; inhibitors should be added immediately before use; the buffer should be kept on ice; and swirling rather than vigorous vortexing is recommended after adding additives.
Additional reagents:
- BCA protein assay reagents
- 50 mM Tris-HCl, pH 8.0
- DTT
- Iodoacetamide
- LysC
- Trypsin
- Formic acid
- TMT reagents, if using multiplexed quantitative proteomics
- IMAC reagents, if performing phosphopeptide enrichment

Ultrasonicator UP200Ht with microtip S26d2 for sample preparation
Equipment for Sonication-Assisted Protein Extraction
Required
Recommended probe selection
For samples around 200–1000 µL, use a small-diameter sonotrode appropriate for direct high-intensity processing of small volumes. Hielscher offers multiple sonotrode diameters, and smaller tip diameters provide higher intensity at the tip.
Practical note: Use the smallest probe that gives efficient mixing without excessive foaming or vessel wall contact.
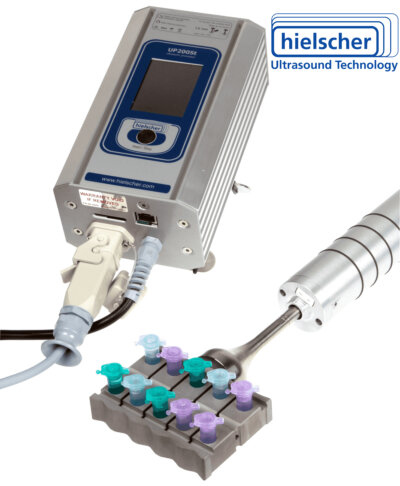
If you work with multiple samples at the same time, you might consider the Multi-Tube Sonicator VialTweeter or the Microplate Sonicator UIP400MTP!
Step-by-Step Instructions: Procedure of Sonication-Assisted Protein Extraction
A. Pre-cool and Prepare
- Chill the centrifuge to 4°C.
- Prepare fresh urea lysis buffer and keep it on ice.
- Set up the UP200Ht or UP200St on a stand inside a sound enclosure.
- Prepare an ice bath large enough to hold sample tubes upright and stable.
Fresh buffer preparation and cold handling are critical.
B. Initial Urea Extraction
- Keep cryopulverized tissue on ice.
- Add 200 µL chilled urea lysis buffer per 50 mg wet tissue.
- Vortex 5–10 s at high speed.
- Incubate 15 min at 4°C.
- Repeat the vortex-plus-incubation step once more.
- Centrifuge at 20,000g for 10 min at 4°C.
- Transfer the lysate/supernatant to a clean low-bind tube.
C. Sonication with using UP200Ht or UP200St
Starting Conditions for Sonication
- Amplitude: start at 20–30%
- Pulse length: 5 s ON
- Cooling: 2 min on ice between pulses
- Number of cycles: 4 cycles
- Total active sonication time: 20 s
- Total process time including cooling: about 8–10 min
Sonication Steps
- Transfer lysate to a tube suited for sonication. Use a narrow, thin-walled tube that allows good heat exchange and safe probe immersion.
- Place the tube in an ice bath. The paper identifies cooling during sonication as critical to prevent heat damage.
- Immerse the probe tip into the sample. Keep the tip submerged sufficiently for stable cavitation, but do not let it touch the tube wall or bottom.
- Run one 5 s pulse at the starting amplitude.
- Immediately return the sample fully to ice for 2 min.
- Repeat until 4 cycles are completed.
- Inspect the lysate after cycle.
- Continue only if needed, using one additional 5 s pulse at a time, always followed by full cooling.
- Stop once the lysate becomes more uniform and less stringy/viscous.
The endpoint is a more translucent lysate that forms drops rather than a continuous viscous flow. - Centrifuge at about 16,000g for 15 min at 4°C.
- Transfer the clarified supernatant to a fresh tube and measure protein concentration.
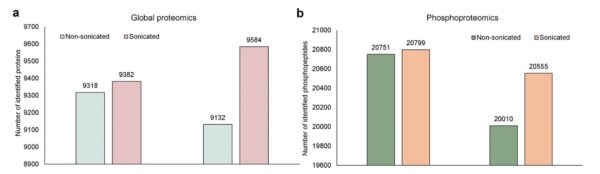
Comparison of global proteomics and phosphoproteomics between sonicated and non-sonicated samples.
a. Number of identified proteins (global proteomics) in all PDX tumor tissues with or without sonication. b. Number of identified phosphopeptides (IMAC enrichment) in all PDX tumor tissues with or without sonication. Only proteins and phosphopeptides with abundance ratio greater than or equal to the 25th percentile were counted. The abundance ratio was calculated between the same types of samples.
Study and graphs: ©Li et al., 2025
Downstream Digestion and Analysis
After sonication and clarification, proceed as described in the original CPTAC-style workflow:
- Dilute lysate 1:3 (v/v) with 50 mM Tris-HCl pH 8.0 to reduce urea to <2 M.
- Add LysC at 1 mAU per 50 µg protein and incubate 2 h at 25°C.
- Add trypsin at 1:49 enzyme:substrate (w/w) and digest overnight at 25°C.
- Quench with formic acid to 1% final.
- Continue to desalting, TMT labeling, fractionation, phosphopeptide enrichment, and LC-MS/MS as required.
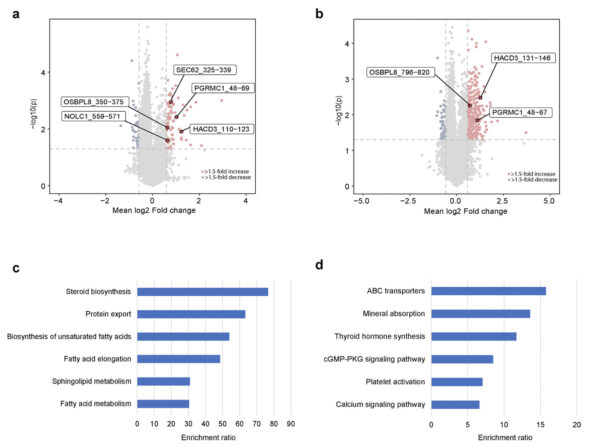
Differential expression of phosphopeptides from TMT-labeled MS.
a. The basal subtype, highlighting some human phosphopeptides (protein-sequence begin and end) upregulated in sonicated samples compared with nonsonicated samples. b. The luminal subtype, highlighting some human phosphopeptides (protein_sequence begin and end) upregulated in sonicated samples compared with nonsonicated samples. c. Enriched KEGG pathways based on the proteins of upregulated phosphopeptides in sonicated basal subtype of tumors. d. Enriched KEGG pathways based on the proteins of upregulated phosphopeptides in sonicated luminal subtype of tumors.
Study and graphs: ©Li et al., 2025
Design, Manufacturing and Consulting – Quality Made in Germany
Hielscher ultrasonicators are well-known for their highest quality and design standards. Robustness and easy operation allow the smooth integration of our ultrasonicators into industrial facilities. Rough conditions and demanding environments are easily handled by Hielscher ultrasonicators.
Hielscher Ultrasonics is an ISO certified company and put special emphasis on high-performance ultrasonicators featuring state-of-the-art technology and user-friendliness. Of course, Hielscher ultrasonicators are CE compliant and meet the requirements of UL, CSA and RoHs.

Multi-well plate Sonicator UIP400MTP for high-throughput protein extraction from samples
Literature / References
- Li Q.K.; Lih T.M.; Clark D.J.; Chen L.; Schnaubelt M.; Zhang H. (2025): Sonication-assisted protein extraction improves proteomic detection of membrane-bound and DNA-binding proteins from tumor tissues. Nature Protocols 2025 Aug;20(8):2083-2099.
- Stadlmann, J., Taubenschmid, J., Wenzel, D. et al. (2017): Comparative glycoproteomics of stem cells identifies new players in ricin toxicity. Nature 549, 2017. 538–542
- Jakob, S., Steinchen, W., Hanßmann, J. et al. (2024): The virulence regulator VirB from Shigella flexneri uses a CTP-dependent switch mechanism to activate gene expression. Nature Communications 15, 318 (2024).
- Jorge S., Capelo J.L., LaFramboise W., Dhir R., Lodeiro C., Santos H.M. (2019): Development of a Robust Ultrasonic-Based Sample Treatment To Unravel the Proteome of OCT-Embedded Solid Tumor Biopsies. Journal of Proteome Research 2019 Jul 5;18(7):2979-2986.
- Mertins P, Tang LC, Krug K, et al. (2018): Reproducible workflow for multiplexed deep-scale proteome and phosphoproteome analysis of tumor tissues by liquid chromatography-mass spectrometry. (Initial CPTAC protocol). Nature Protocols 7, 2018. 1632-1661.
Frequently Asked Questions
What is the Difference between Protein Extraction from Plant Tissue vs Mammalian Tissue?
Protein extraction from plant tissue is usually more difficult than from mammalian tissue because plant cells have a rigid cellulose-rich cell wall, abundant polysaccharides, phenolic compounds, pigments, and active endogenous enzymes that can interfere with protein solubilization, contaminate extracts, or promote protein degradation. By contrast, mammalian tissue lacks a cell wall and is generally easier to disrupt chemically, although it often contains more lipids, extracellular matrix, and proteases that can still complicate extraction. As a result, plant protocols typically require stronger mechanical disruption, low-temperature grinding, and additives such as PVPP, reducing agents, or phenol-based cleanup, whereas mammalian tissue is more often handled successfully with detergent-based buffers such as RIPA or urea-based lysis systems. Both, plant and mammalian tissue, can be efficiently lysed by sonication!
How to Improve Protein Extraction from Tissue using RIPA Buffer?
Protein extraction from tissue with RIPA buffer can be improved by optimizing both disruption and protection of the sample. The most important measures are to keep the tissue and buffer cold, use freshly prepared protease and phosphatase inhibitors, mince or pulverize the tissue thoroughly before lysis, maintain an appropriate buffer-to-tissue ratio, and add a mechanical disruption step such as homogenization or brief probe sonication to break up dense tissue and reduce viscosity. Extraction efficiency also improves when lysates are incubated on ice with intermittent mixing, followed by high-speed refrigerated centrifugation to remove insoluble debris. For fibrous, lipid-rich, or highly structured tissues, repeating the extraction once or increasing detergent exposure time can improve recovery, but excessive sonication or prolonged room-temperature handling should be avoided because they can denature proteins or increase proteolysis.
What Safety Requirements Should be Maintained during Ultrasonic Protein Extraction?
Perform probe sonication in a sound protection box or with suitable hearing protection. The source protocol specifically warns about harmful audible noise generated during sonication.
Wear eye protection, lab coat, and gloves.
Keep the probe away from skin and never sonicate an open vessel without splash protection.
Handle PMSF and other inhibitors according to local chemical safety rules.
Keep samples on ice to reduce heat-induced protein damage; the paper repeatedly identifies temperature control as critical.

The VialTweeter Sonicator accelerates and improves protein extraction in proteomics

Hielscher Ultrasonics manufactures high-performance ultrasonic homogenizers from lab to industrial size.
