Chromatin Shearing with Sonication
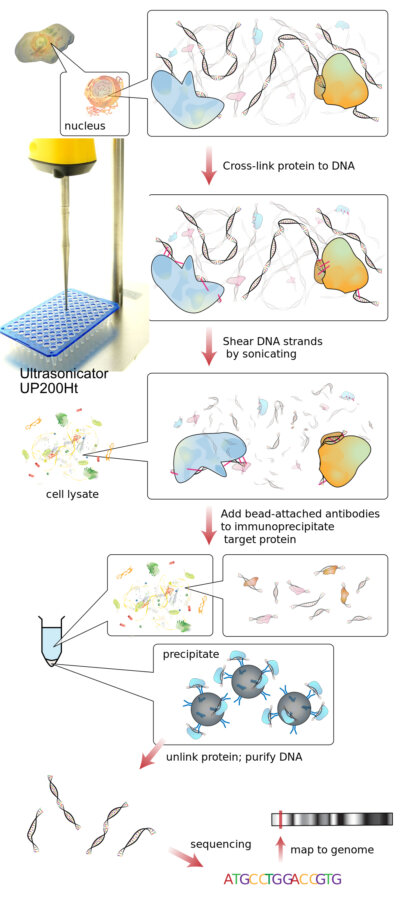
Chromatin shearing is a critical step in many epigenetics and molecular biology workflows, particularly in chromatin immunoprecipitation (ChIP), ChIP-seq, and related assays. The goal is to fragment chromatin into reproducible DNA-protein complexes while preserving epitope integrity and minimizing sample loss. Among the available methods, ultrasonic chromatin fragmentation has become a widely used approach because it provides reliable, reagent-free fragmentation with excellent reproducibility.
What Should I Consider When Shearing Chromatin?
Efficient chromatin shearing requires careful control of experimental parameters. Improper fragmentation can compromise downstream ChIP experiments by generating fragments that are either too large, overly degraded, or inconsistent between samples.
One of the most important factors is the desired fragment size distribution. For most ChIP and ChIP-seq applications, chromatin fragments between 100 and 600 base pairs are optimal. This size range enables efficient immunoprecipitation while providing sufficient resolution for genomic mapping.
 Another key factor is the crosslinking efficiency prior to sonication. Most ChIP workflows involve formaldehyde fixation to stabilize protein-DNA interactions. However, excessive crosslinking can make chromatin more resistant to fragmentation, requiring longer sonication times and potentially increasing heat exposure.
Another key factor is the crosslinking efficiency prior to sonication. Most ChIP workflows involve formaldehyde fixation to stabilize protein-DNA interactions. However, excessive crosslinking can make chromatin more resistant to fragmentation, requiring longer sonication times and potentially increasing heat exposure.
Temperature control is also crucial. Sonication generates localized energy that can raise sample temperature. Elevated temperatures may damage DNA or denature proteins, affecting antibody recognition during ChIP. Many researchers therefore perform pulsed sonication cycles combined with cooling intervals to maintain sample stability.
Sample concentration and volume also influence fragmentation efficiency. Highly concentrated chromatin suspensions may require longer sonication times, while small sample volumes demand precise energy delivery to prevent over-processing.
Finally, the choice of sonication device strongly influences experimental reproducibility. Devices designed for chromatin shearing typically provide controlled ultrasonic energy and standardized sample handling, enabling consistent fragmentation across multiple samples.
Which Sonicator Should I Choose for Chromatin Shearing?
Different laboratory workflows require different sonication configurations. The optimal system depends largely on sample throughput, volume, and experimental format.
Probe-Type Sonicator
A probe-type sonicator delivers ultrasonic energy directly into the sample through a titanium probe. This configuration provides very high energy density and is therefore suitable for robust chromatin disruption in individual samples.
Probe sonicators are particularly useful for:
- Small-to-medium sample numbers
- Difficult-to-fragment chromatin
- Flexible experimental protocols
Multi-Tube Sonicator – VialTweeter
For laboratories processing multiple samples simultaneously, the VialTweeter multi-tube sonicator provides a highly reproducible solution. The system transmits ultrasonic energy indirectly through the vial holder, allowing several sealed tubes to be fragmented under identical conditions.
This configuration offers important advantages:
- Parallel chromatin shearing of multiple samples
- Elimination of probe contamination
- High reproducibility between tubes
- Simplified workflow for ChIP sample preparation
Such multi-tube systems are well suited for routine ChIP experiments and medium-throughput studies.
Microplate Sonicator – UIP400MTP
High-throughput epigenetics studies increasingly rely on microplate-based sample processing. The UIP400MTP microplate sonicator is designed to fragment chromatin directly in standard microplates without transferring samples.
This approach enables:
- Simultaneous processing of dozens or hundreds of samples
- Automation-friendly workflows
- Uniform ultrasonic energy distribution across wells
- Significant reduction in sample handling steps
For large ChIP-seq screening projects or high-throughput epigenetic studies, microplate sonication provides exceptional scalability and efficiency. The multi-well plate sonicator UIP400MTP is well-suited for integration into liquid handling systems and automated lab workflows.
Why Choosing Sonication over Other Chromatin Shearing Techniques?
Compared with enzymatic approaches, sonication for ChIP provides unbiased fragmentation, since the process does not depend on sequence-specific enzyme activity. This is particularly important for genome-wide epigenetic studies, where uniform coverage is essential.
Another major advantage is scalability. Ultrasonic systems can accommodate single samples, multiple tubes, or entire microplates, allowing laboratories to select the most suitable configuration for their experimental throughput.
Finally, sonication provides excellent control over fragmentation parameters. By adjusting pulse cycles, duration, and power levels, researchers can reliably achieve the desired fragment size distribution.
Comparison of Chromatin Shearing Techniques
| Chromatin Shearing Method | Principle | Advantages | Limitations |
| Sonication | High-frequency acoustic energy mechanically fragments chromatin. | Reagent-free fragmentation, highly reproducible results, tunable fragment size distribution, compatible with crosslinked chromatin, scalable from single tubes to multi-sample and microplate formats. | Requires sonication equipment and optimization of sonication parameters. |
| Enzymatic Digestion (MNase) | Micrococcal nuclease digests DNA between nucleosomes. | Gentle fragmentation and useful for native chromatin analysis. | Enzyme bias, sequence preference, digestion difficult to control, potential variability between experiments. |
| Mechanical Shearing (Needle / Syringe) | Chromatin is disrupted through repeated physical force. | Simple method that requires minimal equipment. | Poor reproducibility, limited control over fragment size, labor-intensive for multiple samples. |
| Nebulization | Compressed air forces DNA through small orifices causing fragmentation. | Rapid fragmentation process. | Possible sample loss, limited scalability, requires specialized equipment. |
How Do I Quantify and Qualify Chromatin Yield after Ultrasonic Fragmentation?
 After sonication for ChIP, researchers must evaluate both the quantity and quality of fragmented chromatin. This verification step ensures that chromatin fragmentation meets the requirements of downstream applications such as ChIP-qPCR or ChIP-seq.
After sonication for ChIP, researchers must evaluate both the quantity and quality of fragmented chromatin. This verification step ensures that chromatin fragmentation meets the requirements of downstream applications such as ChIP-qPCR or ChIP-seq.
Quantification typically begins with measuring DNA concentration. Spectrophotometric methods such as Nanodrop analysis or fluorometric assays like Qubit DNA quantification provide reliable estimates of chromatin yield following decrosslinking and purification.
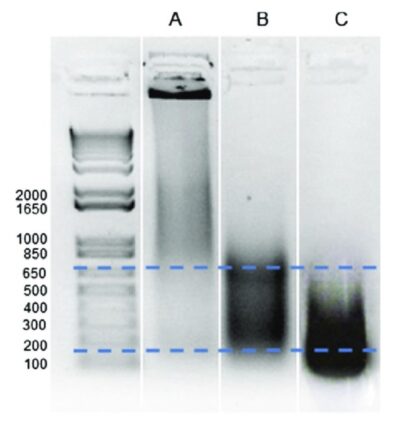
However, DNA concentration alone does not reveal whether fragmentation was successful. Researchers therefore assess fragment size distribution using electrophoretic techniques. Agarose gel electrophoresis remains a commonly used approach for visualizing DNA fragments and verifying that the majority fall within the target size range.
More advanced laboratories often use capillary electrophoresis systems, such as the Agilent Bioanalyzer or TapeStation. These platforms provide precise size distribution profiles and allow researchers to detect over-fragmentation or incomplete shearing.
When evaluating chromatin quality after ultrasonic fragmentation, researchers typically confirm:
- The majority of DNA fragments fall within the 100–600 bp range
- Fragment distribution is consistent across replicate samples
- DNA degradation is minimal
- Total chromatin yield is sufficient for the planned ChIP assay
Proper quality control ensures that the ultrasonic chromatin shearing step produces reproducible and biologically meaningful results.
Conclusion: Ultrasonic Chromatin Shearing for Reliable Research
Reliable chromatin shearing is fundamental for successful ChIP and epigenetics research. Ultrasonic fragmentation offers a powerful solution because it enables precise, reproducible, and reagent-free chromatin disruption across a wide range of experimental formats.
By carefully optimizing sonication parameters, verifying fragment size distribution, and selecting the appropriate sonication system – whether a probe-type sonicator, multi-tube VialTweeter, or the high-throughput UIP400MTP microplate sonicator – researchers can achieve consistent chromatin fragmentation that supports high-quality ChIP and ChIP-seq results.
As epigenetics research continues to expand toward higher throughput and greater experimental reproducibility, ultrasonic chromatin shearing remains one of the most versatile and reliable methods available for modern molecular biology laboratories.
Design, Manufacturing and Consulting – Quality Made in Germany
Hielscher ultrasonicators are well-known for their highest quality and design standards. Robustness and easy operation allow the smooth integration of our ultrasonicators into industrial facilities. Rough conditions and demanding environments are easily handled by Hielscher ultrasonicators.
Hielscher Ultrasonics is an ISO certified company and put special emphasis on high-performance ultrasonicators featuring state-of-the-art technology and user-friendliness. Of course, Hielscher ultrasonicators are CE compliant and meet the requirements of UL, CSA and RoHs.
Literature / References
- Dreyer J., Ricci G., van den Berg J., Bhardwaj V., Funk J., Armstrong C., van Batenburg V., Sine C., VanInsberghe M.A., Marsman R., Mandemaker I.K., di Sanzo S., Costantini J., Manzo S.G., Biran A., Burny C., Völker-Albert M., Groth A., Spencer S.L., van Oudenaarden A., Mattiroli F. (2024): Acute multi-level response to defective de novo chromatin assembly in S-phase. Molecular Cell 2024.
- Mittal, N., Guimaraes, J.C., Gross, T. et al. (2017): The Gcn4 transcription factor reduces protein synthesis capacity and extends yeast lifespan. Nat Commun 8, 457 (2017).
- Shih H.-T., Chen W.-Y., Liu K.-Y., Shih Z.-S., Chen Y.-J., Hsieh P.-C., et al. (2016): dBRWD3 Regulates Tissue Overgrowth and Ectopic Gene Expression Caused by Polycomb Group Mutations. PLoS Genetics 12(9): e1006262.
- José A. Jarillo, Dorota N. Komar, and Manuel Piñeiro (2018): The Use of the Chromatin Immunoprecipitation Technique for In Vivo Identification of Plant Protein–DNA Interactions. Chapter in book: Luis Oñate-Sánchez (ed.), Two-Hybrid Systems: Methods and Protocols, Methods in Molecular Biology, vol. 1794, 2018.
- Einig, E., Jin, C., Andrioletti, V. et al. (2023): RNAPII-dependent ATM signaling at collisions with replication forks. Nature Communications 14, 5147 (2023)
Frequently Asked Questions
What is Chromatin?
Chromatin is the structural complex of DNA and associated proteins that organizes genetic material within the nucleus of eukaryotic cells. The primary proteins in chromatin are histones, around which DNA is wrapped to form nucleosomes. This organization compacts DNA while simultaneously regulating access to genetic information for processes such as transcription, replication, and DNA repair.
What are the Types of Chromatin?
Chromatin is generally classified into two main forms: euchromatin and heterochromatin. Euchromatin is loosely packed and transcriptionally active, allowing genes to be readily accessed by the transcriptional machinery. Heterochromatin is more densely packed and transcriptionally inactive, typically containing repetitive DNA sequences or genes that are silenced. Heterochromatin can be further divided into constitutive heterochromatin, which remains permanently condensed, and facultative heterochromatin, which can switch between active and inactive states depending on cellular conditions.
What is Crosslinking?
Crosslinking is a biochemical process used to stabilize interactions between biomolecules by forming covalent bonds between them. In chromatin research, crosslinking is commonly used to preserve protein–DNA interactions within chromatin prior to analysis. Chemical agents such as formaldehyde are typically used to create reversible covalent links between DNA and associated proteins, effectively “freezing” molecular interactions at a specific moment in time. This stabilization allows chromatin complexes to be fragmented and processed without losing the native associations between DNA and regulatory proteins, which is essential for techniques such as chromatin immunoprecipitation (ChIP).
What is ChIP?
Chromatin immunoprecipitation (ChIP) is a molecular biology technique used to investigate interactions between proteins and DNA within chromatin. In this method, DNA-protein complexes are first stabilized, typically by crosslinking, and chromatin is then fragmented. Antibodies specific to a target protein are used to immunoprecipitate the protein-DNA complexes, allowing the associated DNA sequences to be isolated and analyzed.
What is ChIP used for?
ChIP is used to identify genomic regions bound by specific DNA-associated proteins such as transcription factors, histone modifications, or chromatin-associated regulatory proteins. The technique is widely applied to study gene regulation, epigenetic modifications, transcription factor binding sites, and chromatin structure. When combined with downstream analytical methods such as quantitative PCR (ChIP-qPCR) or high-throughput sequencing (ChIP-seq), it enables genome-wide mapping of protein–DNA interactions.
What are the Types of ChIP?
Several variants of chromatin immunoprecipitation exist depending on the experimental design and downstream analysis. The most common approaches include ChIP-qPCR, which quantifies enrichment of specific genomic regions; ChIP-seq, which uses next-generation sequencing to map protein-DNA interactions across the genome; and ChIP-chip, which combines ChIP with DNA microarray analysis. Additional variants such as native ChIP (N-ChIP), which analyzes non-crosslinked chromatin, and crosslinked ChIP (X-ChIP), which uses chemical crosslinking to stabilize protein-DNA interactions, are also widely used depending on the biological question being investigated.

Hielscher Ultrasonics manufactures high-performance ultrasonic homogenizers from lab to industrial size.